Complexity. Beauty. Mystery.
The world of single-celled organisms is mostly still to be discovered, says Dr Ross Waller.
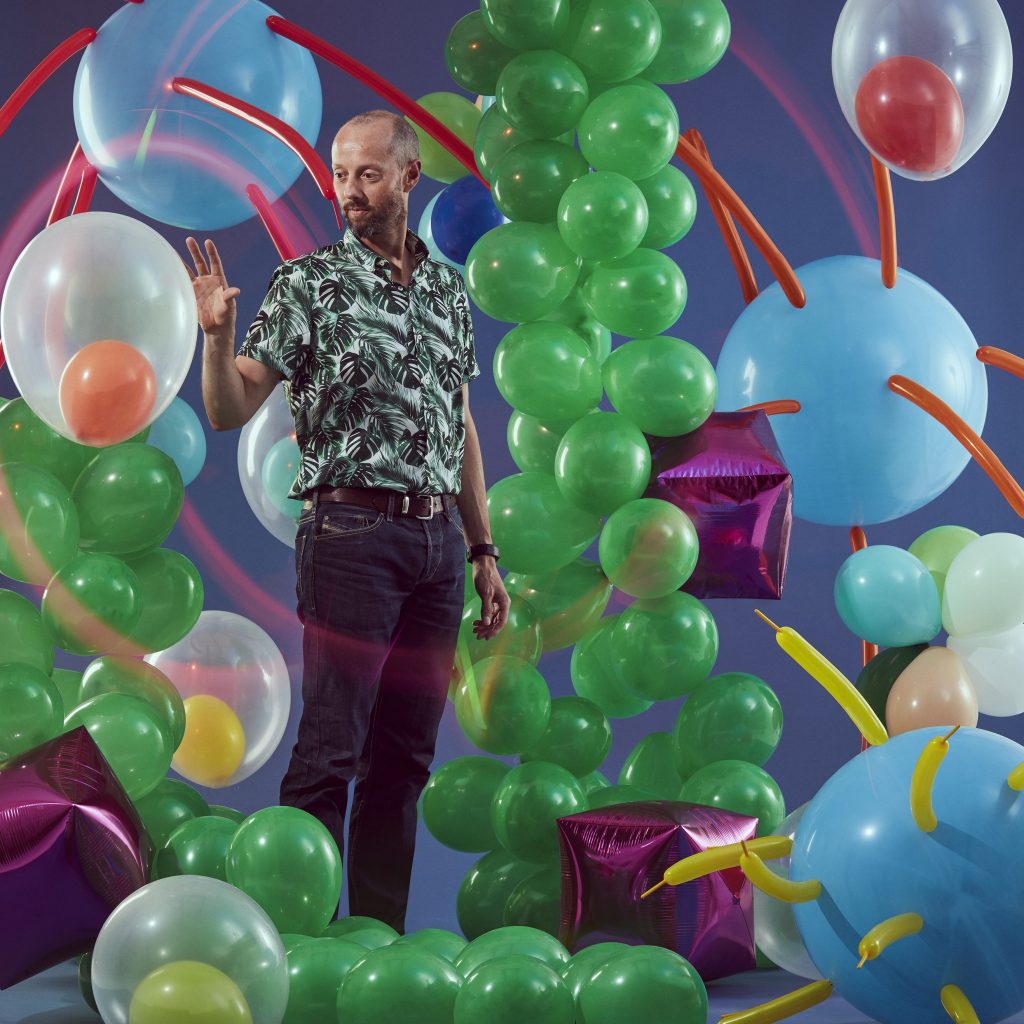

You can see them from space, spread out across the oceans in great swathes of blue and green. You can feel them in the ground beneath your feet, built up over millions of years. They live inside most animals on the planet, including ourselves. They’re responsible for some of the most dangerous diseases known to humanity, but they are also a vital part of the systems and forces that keep the Earth alive. Yet we barely acknowledge their existence.
Welcome to the universe of single-celled organisms. It’s a strange and beautiful place of complexity, beauty and mystery – because although the idea of a single cell might sound simple, it is anything but. It’s what drew Wellcome Trust investigator and Reader in Cell Biology and Evolution Dr Ross Waller to study them in the first place – and what keeps him endlessly fascinated. “These are not just blobs that bounce around in the ocean, or the soil, or your body,” he says. “They have complex, successful behaviours. They are very active. They have purpose – for example, at least half the planet’s photosynthesis occurs at the single-cell level, particularly in the ocean. And they are often stunningly beautiful.”
They also represent the basic units of life from which all multicellular organisms, such as humans, plants and animals, have evolved. “In terms of the fundamental diversity of life, the majority of that is at the single-cell level,” Waller says. “At the level of the cell, all animals have a lot in common. If you want to understand life processes, you get a limited view if you look at just one group of organisms such as animals. “But look at single-celled organisms and you see how differently the challenges or objectives of life have been solved – which tells us a lot more about these processes.”
Waller’s lab, down the twisting corridors of the Hopkins Building, studies two very different families of single-celled organisms. One is deadly. The apicomplexans family of pathogens includes the malaria parasite Plasmodium, (which causes 400,000 deaths and 200 million infections every year), toxoplasmosis, (which infects a third of all humans) and cryptosporidiosis (which contributes to 800,000 infant deaths annually). Apicomplexans also infects every known species of animal. “We primarily work on ones that live in humans, because of the medical relevance,” says Waller. “It’s work on a fundamental level. We want to learn more about how the cell has adapted, including how it develops innovations to interact with humans and exploit them. These are complex parasites. They live and grow inside our cells. It’s a very intimate relationship, and they have become very good at it.”
Know your enemy, that’s the old saying. And we don’t. We’ve only scratched the surface of understanding single-celled organisms.
The malaria parasite, in particular, is a terrifying foe. It has learned to avoid the human immune system when it goes on the attack: disguising itself from our defensive cells, or hiding in a place where our bodies can’t attack them. “Know your enemy, that’s the old saying,” says Waller. “And we don’t. We’ve only scratched the surface of understanding single-celled organisms. Even with the better understood ones, such as the malaria parasite, we still only understand a fraction of the cell’s complexity. If we don’t know or understand our enemy, we can’t defend ourselves against them.”
But the other group, dinoflagellates, seems to have a very different purpose: they help to keep our planet alive. You are unlikely to be familiar with them. Science isn’t particularly familiar with them, either – and that, Waller says, is why they’re so fascinating to study.
Dinoflagellates are named for their shape – the Greek dinos (whirling) and the Latin flagellum (whip, scourge) – rather than any potentially painful properties. They spend most of their time swimming around the ocean, producing food from sunlight and in turn feeding the oceans, or in a deep relationship with coral reefs. This relationship is complex and important: the dinoflagellate feeds the coral, while the coral provides it essential nutrients. When that relationship breaks down – from causes that also aren’t fully understood, but which include rising ocean temperatures – the coral bleaches and dies.
Coral
Consider the consequences of widespread coral loss and you’ll start to see how vital it is to understand dinoflagellates, says Waller. “We don’t understand that relationship well enough to predict what will happen – let alone any possibility of managing it. We need to get far more of an insight into the diversity of these single-celled organisms, to work out how they fit in these complex biological networks.”
But so far, these tiny organisms remain an enigma. One well-accepted way of studying a single-celled organism is by assembling a list of all of its molecular components. This can be done by sequencing the genome – the ‘parts list’. But a list of parts on its own is not much use: for Plasmodium and Toxoplasma we have the parts list, but don’t know what 60 per cent of the parts are – or what they do. It’s like trying to read a book in a language you’ve only just started learning, says Waller. Hard, but not impossible.
Our biology isn’t separate from the biology of the planet. We need to understand how we are interacting with it and how it is interacting with us
“We want to put all that information in context – how is it driving the complexity and architecture of the cells. Think of the exploded diagrams common to engineering (or, indeed, flat-pack furniture), where every part is carefully drawn and which shows you how it fits together. “We’re using a method, developed in the Department by Professor Kathryn Lilley, which overlays the genome-level information of parts on to the cell, enabling us to locate those various parts. “We can then look at the function of each of these parts by using genetic tools such as Crispr to take them out, to turn them on or off, to modify their behaviours.”
A new parts list
But if Plasmodium and Toxoplasma are Madame Bovary to a French A-level student, then dinoflagellates are Linear A. Crispr doesn’t work on them, says Waller, because they’ve radically changed the way they store their genetic information (i.e. the parts list). Most complex organisms, including us, fall into a class known as eukaryotes. These cells have developed a way of managing their genomes in a nucleus, using proteins called histones. The genome’s chromosomes are organised on histones, like scrolls in a library. This efficient system for organising their genetic information – evolved over two billion years of life on Earth – enables them to be more complex. All eukaryotes use this system. Except the dinoflagellates.
“In biology, you can almost never say never. There’s almost always an exception,” says Waller, cheerfully. “And dinoflagellates are one of the most remarkable exceptions. They have got rid of the histone system for organising their data. “And that’s remarkable, because their genome is up to a hundredfold bigger than the human genome. So if you think of the nucleus as about the size of a basketball, in a human nucleus the DNA would be the equivalent of 30 miles of string, all stored inside that basketball. At that scale, dinoflagellates have about 2,500 miles of string to store – and stored so that it can get to any part of the string whenever it needs to.”
That’s a huge paradox: why would a single cell make its genetic information much bigger than complex things like humans? And how does it access this information? They are compacting it somehow, Waller says, in a way that still keeps it accessible. “And this is now an opportunity to learn about a fundamental cell process, or life process, of organising DNA in a complex way that’s different from what the rest of complex cellular life does.”
From an early age, Waller says, he was fascinated by how systems worked. Initially enrolled to study engineering, he found his first year uninspiring and, by his third year, he had changed to biology. It’s not such a leap, he says, if you think of cells as complex machines built from components, each with its place in the system. “And I was worried about where we’re going as a society, environmentally speaking. As a child in the 90s, when the greenhouse effect and acid rain were very much in the media, I recognised the impact of human activity on these basic processes. I worried about where we were heading. That hasn’t changed.”
Following his degree, he spent a year in a research lab, then signed up for a job on an oceanographic research vessel carrying out phytoplankton sampling, then travelled around the world before returning to research and his PhD at the University of Melbourne, investigating the genetics of Plasmodium. Waller came to Cambridge with his scientist partner who was offered the chance to set up her own lab in nearby Norwich. “And there was an opportunity here. It was just fortuitous. But it’s been a stimulating experience to make new connections. In fact, a lot of the work that we’re doing now is based on new interactions that have been made possible by other researchers at Cambridge.”
His focus now is to keep asking questions and going where they take him, with eyes wide open. “I think we tend to be a bit too blinkered, where we think that the most important thing is to understand human biology,” he says. “Our biology isn’t separate from the biology of the planet. We need to understand how we are interacting with it and how it is interacting with us, as it creates processes that are essential for our lives, and for the planet.”
Find out more about The Waller Lab